import numpy as np
import matplotlib.pyplot as plt
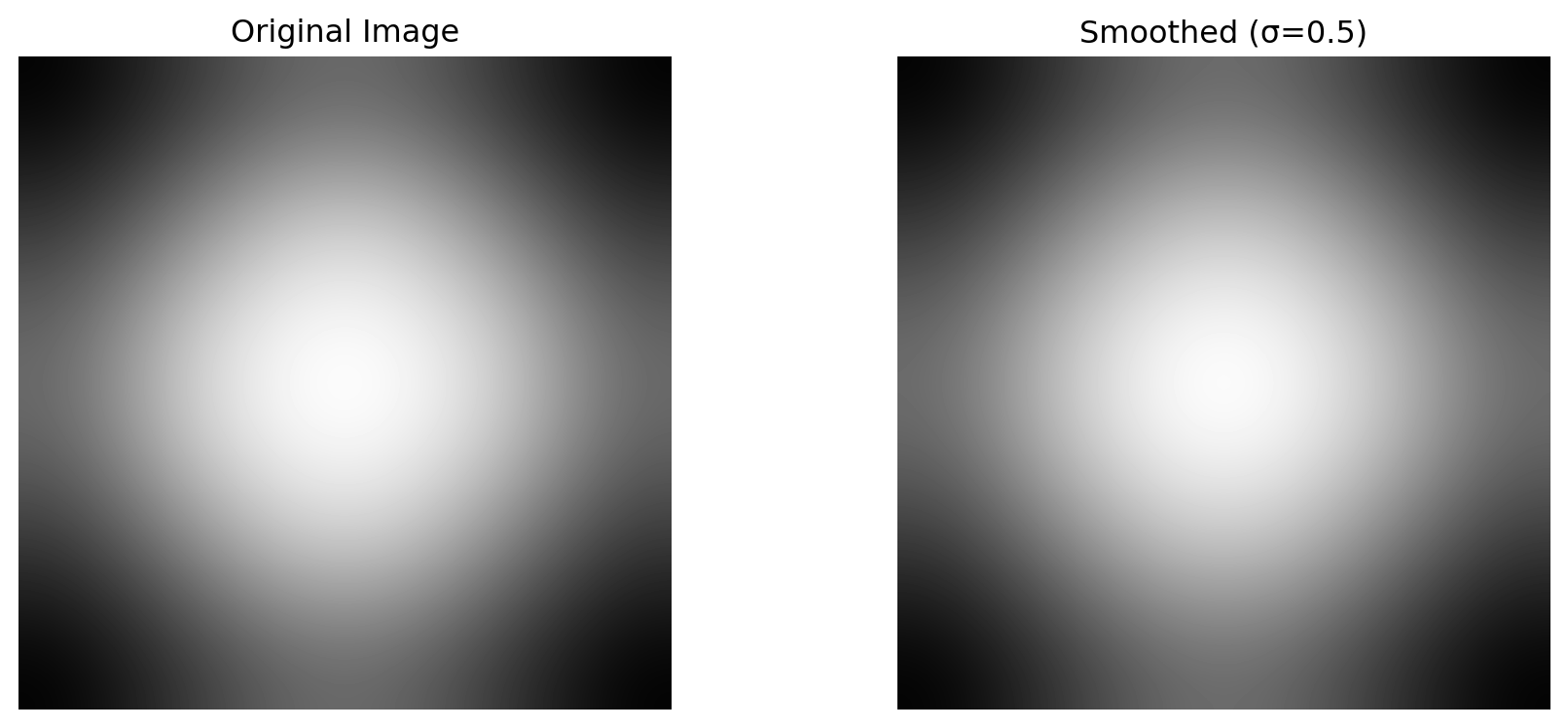
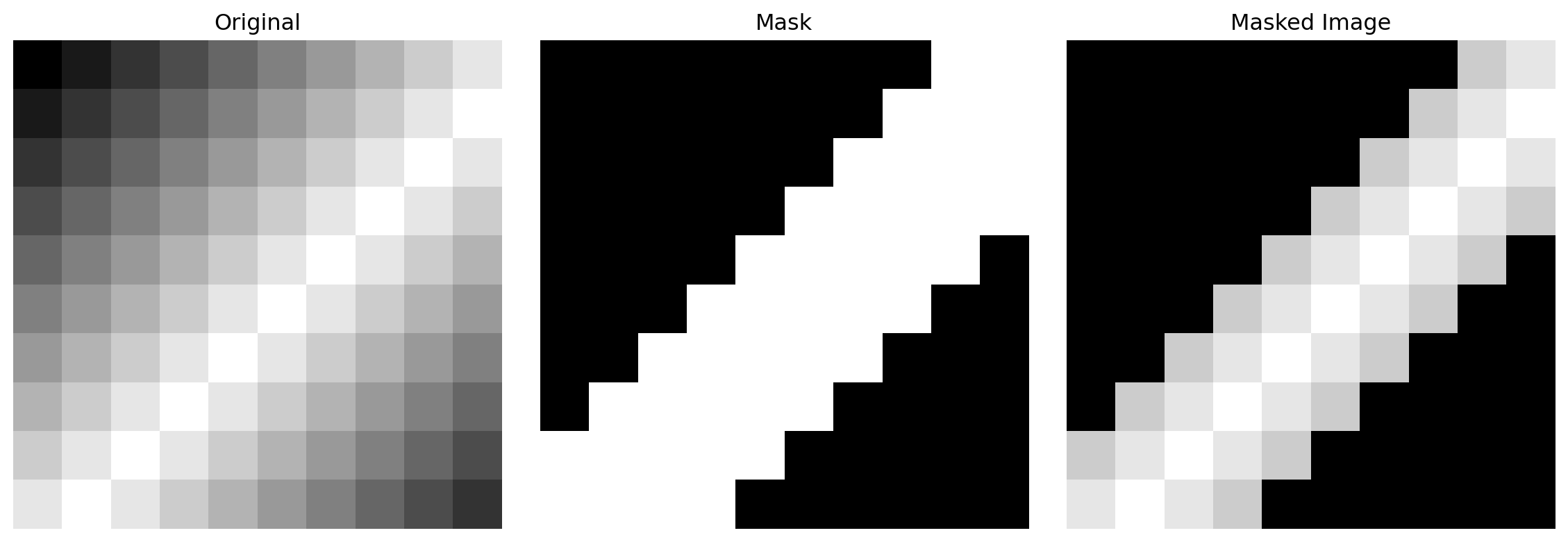
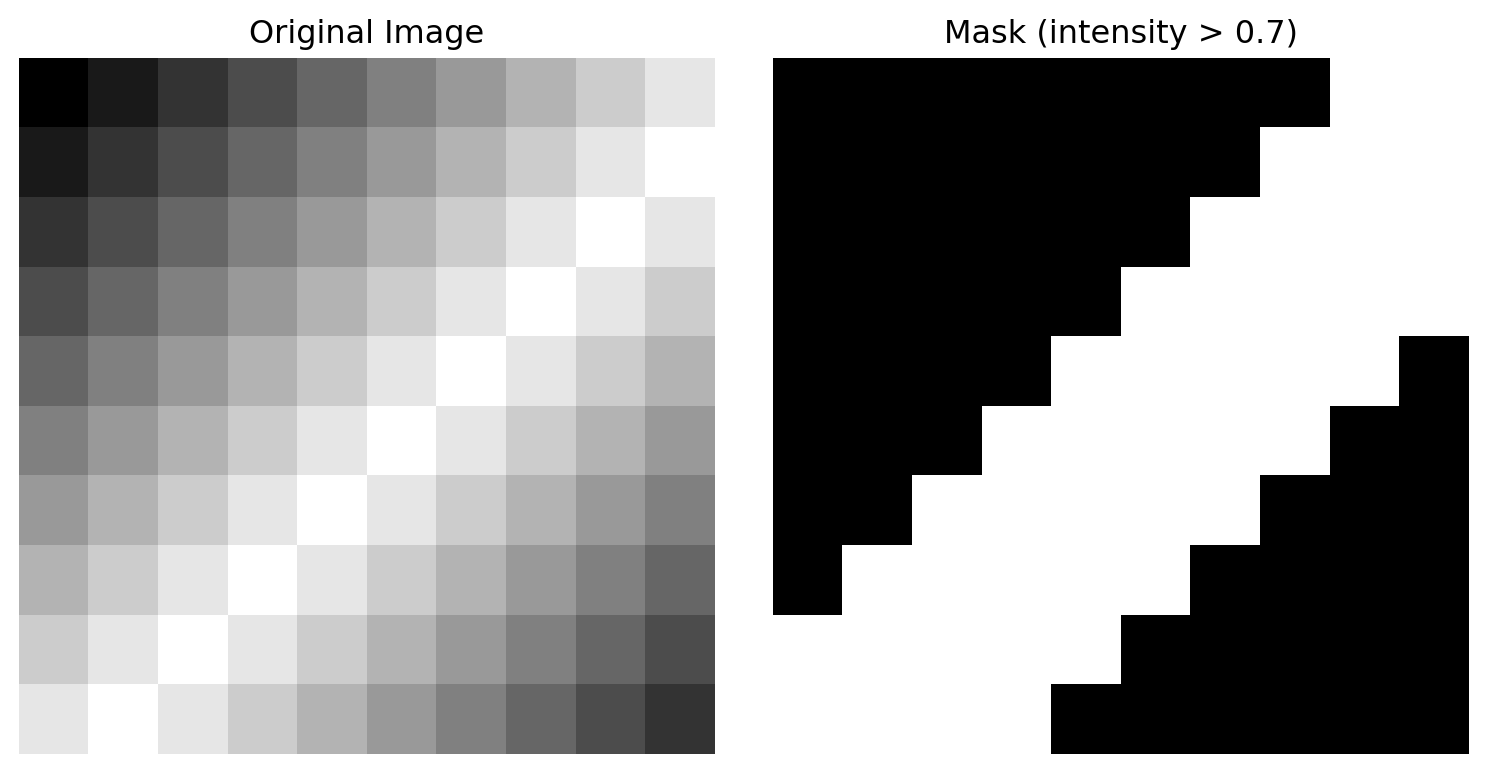
image = np.array([
[0.0, 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9],
[0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0],
[0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0, 0.9],
[0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0, 0.9, 0.8],
[0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0, 0.9, 0.8, 0.7],
[0.5, 0.6, 0.7, 0.8, 0.9, 1.0, 0.9, 0.8, 0.7, 0.6],
[0.6, 0.7, 0.8, 0.9, 1.0, 0.9, 0.8, 0.7, 0.6, 0.5],
[0.7, 0.8, 0.9, 1.0, 0.9, 0.8, 0.7, 0.6, 0.5, 0.4],
[0.8, 0.9, 1.0, 0.9, 0.8, 0.7, 0.6, 0.5, 0.4, 0.3],
[0.9, 1.0, 0.9, 0.8, 0.7, 0.6, 0.5, 0.4, 0.3, 0.2],
], dtype=np.float32)
print(f"Image shape: {image.shape}")
print(f"Value range: [{image.min():.1f}, {image.max():.1f}]")