import numpy as np
import matplotlib.pyplot as plt
from scipy import signal
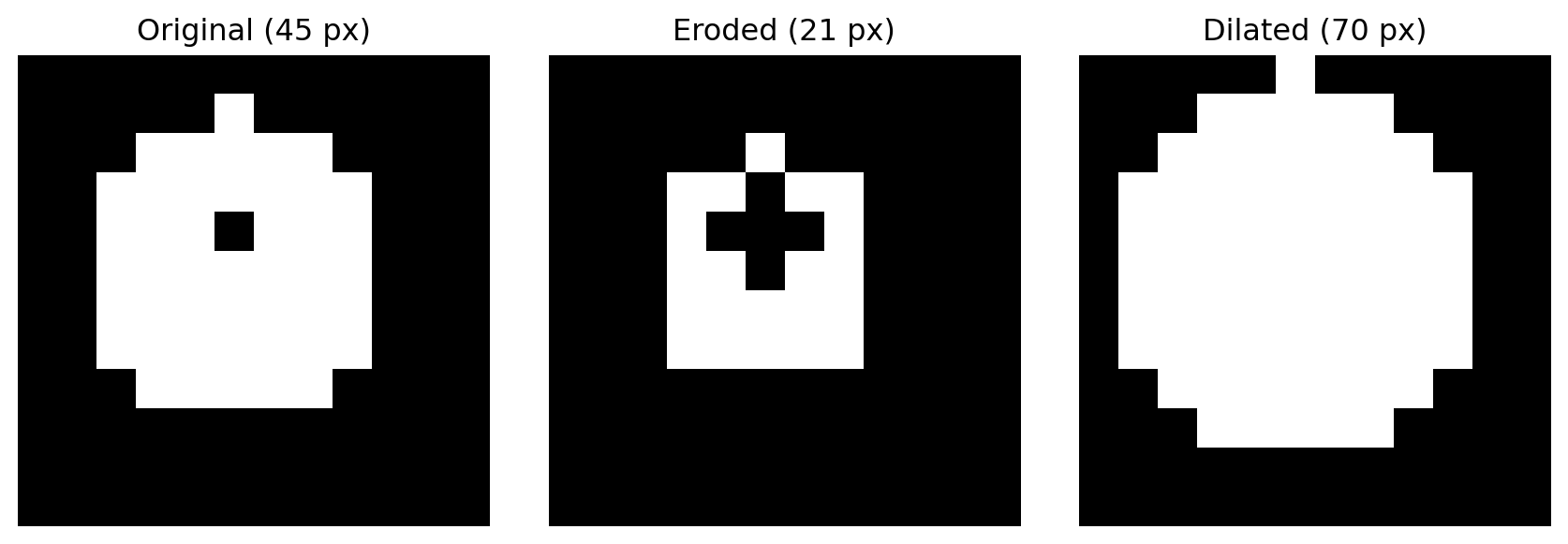
# Create synthetic image: nucleus (low intensity) surrounded by cytoplasm (high intensity)
# White 2-pixel border represents the image background/boundary
nucleus_cytoplasm = np.array([
[1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0],
[1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0],
[1.0, 1.0, 0.4, 0.4, 0.4, 0.4, 0.4, 0.4, 1.0, 1.0],
[1.0, 1.0, 0.4, 0.2, 0.2, 0.2, 0.2, 0.4, 1.0, 1.0],
[1.0, 1.0, 0.4, 0.2, 0.2, 0.2, 0.2, 0.4, 1.0, 1.0],
[1.0, 1.0, 0.4, 0.2, 0.2, 0.2, 0.2, 0.4, 1.0, 1.0],
[1.0, 1.0, 0.4, 0.2, 0.2, 0.2, 0.2, 0.4, 1.0, 1.0],
[1.0, 1.0, 0.4, 0.4, 0.4, 0.4, 0.4, 0.4, 1.0, 1.0],
[1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0],
[1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0]
], dtype=np.float32)
print("Original image (nucleus with white 2-pixel border):")
print(nucleus_cytoplasm)
# Define Sobel kernels
S_x = np.array([
[-1, 0, +1],
[-2, 0, +2],
[-1, 0, +1]
], dtype=np.float32)
S_y = np.array([
[-1, -2, -1],
[0, 0, 0],
[+1, +2, +1]
], dtype=np.float32)
print("\nSobel kernel for x-direction (S_x) - vertical edge detection:")
print(S_x)
print("\nSobel kernel for y-direction (S_y) - horizontal edge detection:")
print(S_y)
# Apply Sobel using correlate2d (standard Sobel without kernel flipping)
# Note: scipy.signal.correlate2d does NOT flip the kernel (unlike ndimage.convolve)
# This matches the standard Sobel implementation in skimage.filters.sobel()
G_x = signal.correlate2d(nucleus_cytoplasm, S_x, mode='same', boundary='fill', fillvalue=0.0)
G_y = signal.correlate2d(nucleus_cytoplasm, S_y, mode='same', boundary='fill', fillvalue=0.0)
# Compute gradient magnitude
G_magnitude = np.sqrt(G_x**2 + G_y**2)
# Compute gradient angle (for reference)
G_angle = np.arctan2(G_y, G_x)
print("\nGradient in x-direction (G_x) - vertical edge detection:")
print(np.around(G_x, decimals=2))
print("\nGradient in y-direction (G_y) - horizontal edge detection:")
print(np.around(G_y, decimals=2))
print("\nGradient magnitude (edges highlighted):")
print(np.around(G_magnitude, decimals=2))
# Visualize
fig, axes = plt.subplots(2, 3, figsize=(16, 10))
# Original image
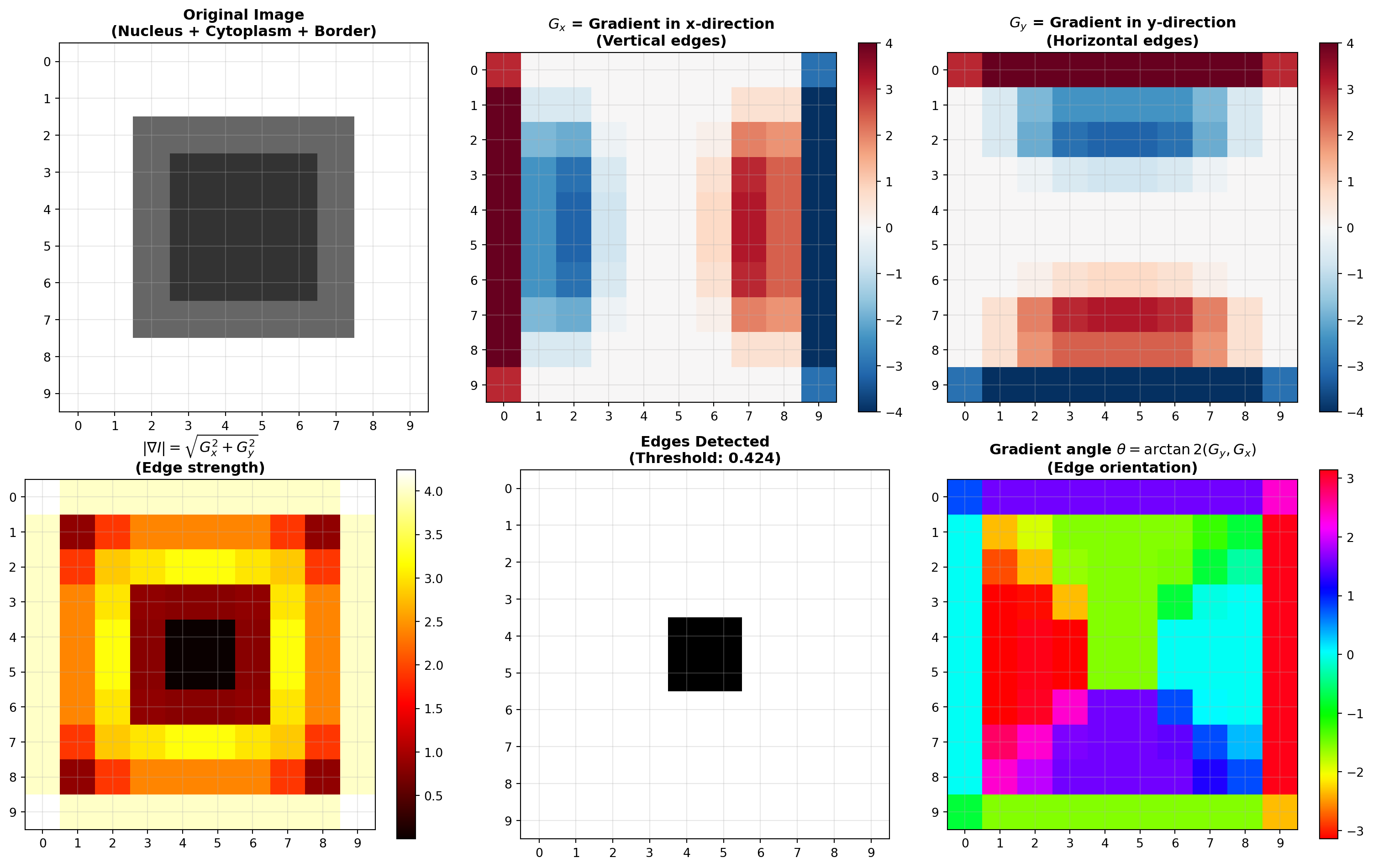
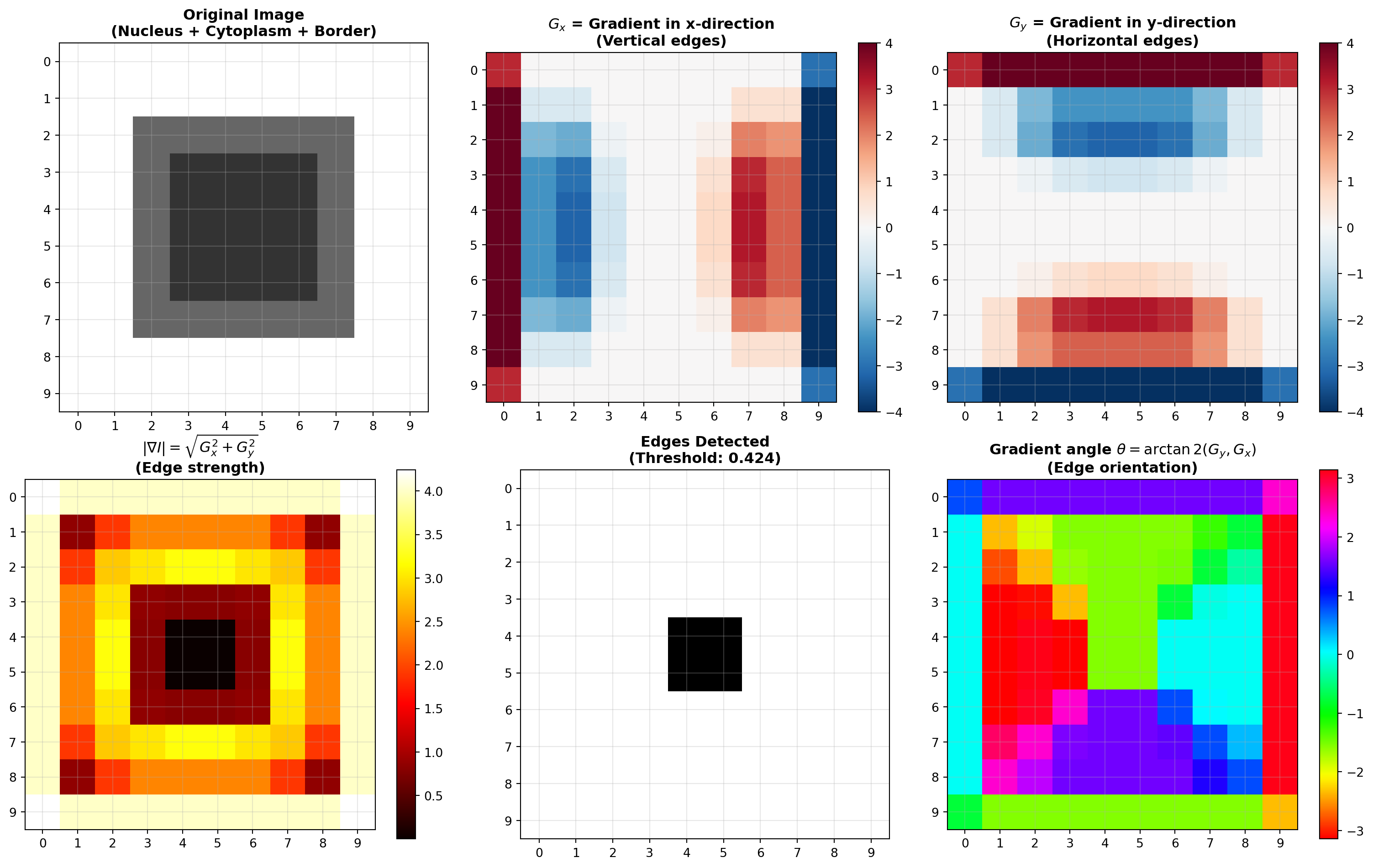
axes[0, 0].imshow(nucleus_cytoplasm, cmap='gray', interpolation='nearest', vmin=0, vmax=1)
axes[0, 0].set_title("Original Image\n(Nucleus + Cytoplasm + Border)", fontsize=12, fontweight='bold')
axes[0, 0].grid(True, alpha=0.3)
axes[0, 0].set_xticks(range(10))
axes[0, 0].set_yticks(range(10))
# Gradient in x-direction
im_gx = axes[0, 1].imshow(G_x, cmap='RdBu_r', interpolation='nearest')
axes[0, 1].set_title(r"$G_x$ = Gradient in x-direction" + "\n(Vertical edges)", fontsize=12, fontweight='bold')
axes[0, 1].grid(True, alpha=0.3)
axes[0, 1].set_xticks(range(10))
axes[0, 1].set_yticks(range(10))
plt.colorbar(im_gx, ax=axes[0, 1])
# Gradient in y-direction
im_gy = axes[0, 2].imshow(G_y, cmap='RdBu_r', interpolation='nearest')
axes[0, 2].set_title(r"$G_y$ = Gradient in y-direction" + "\n(Horizontal edges)", fontsize=12, fontweight='bold')
axes[0, 2].grid(True, alpha=0.3)
axes[0, 2].set_xticks(range(10))
axes[0, 2].set_yticks(range(10))
plt.colorbar(im_gy, ax=axes[0, 2])
# Gradient magnitude
im_mag = axes[1, 0].imshow(G_magnitude, cmap='hot', interpolation='nearest')
axes[1, 0].set_title(r"$|\nabla I| = \sqrt{G_x^2 + G_y^2}$" + "\n(Edge strength)", fontsize=12, fontweight='bold')
axes[1, 0].grid(True, alpha=0.3)
axes[1, 0].set_xticks(range(10))
axes[1, 0].set_yticks(range(10))
plt.colorbar(im_mag, ax=axes[1, 0])
# Gradient magnitude with threshold for edge detection
edge_threshold = 0.1 * G_magnitude.max()
edges_detected = G_magnitude > edge_threshold
axes[1, 1].imshow(edges_detected, cmap='gray', interpolation='nearest')
axes[1, 1].set_title(f"Edges Detected\n(Threshold: {edge_threshold:.3f})", fontsize=12, fontweight='bold')
axes[1, 1].grid(True, alpha=0.3)
axes[1, 1].set_xticks(range(10))
axes[1, 1].set_yticks(range(10))
# Gradient angle
im_angle = axes[1, 2].imshow(G_angle, cmap='hsv', interpolation='nearest')
axes[1, 2].set_title(r"Gradient angle $\theta = \arctan2(G_y, G_x)$" + "\n(Edge orientation)", fontsize=12, fontweight='bold')
axes[1, 2].grid(True, alpha=0.3)
axes[1, 2].set_xticks(range(10))
axes[1, 2].set_yticks(range(10))
plt.colorbar(im_angle, ax=axes[1, 2])
plt.tight_layout()
plt.show()
print("\nEdge Detection Summary:")
print(f" Maximum gradient magnitude: {G_magnitude.max():.4f}")
print(f" Edge threshold (10% of max): {edge_threshold:.4f}")
print(f" Number of edge pixels: {np.sum(edges_detected)}")
print(f" Edge pixels represent: {100 * np.sum(edges_detected) / edges_detected.size:.1f}% of image")